About SWFSbase
DNA-based taxonomic identification relies on comparison to reference databases. While public repositories such as GenBank are invaluable, they are not designed for forensic use and may lack critical metadata or contain misidentified records. In addition, many wildlife forensic laboratories maintain curated internal databases that are not readily accessible to the wider community.
SWFSbase is a secure BLAST server adapted specifically for the wildlife forensic community. It centralises curated reference databases from contributing laboratories and links them to essential forensic metadata, while also incorporating selected publicly available data. SWFSbase strengthens the reliability and transparency of reference data used in wildlife forensic analysis. The platform is hosted on SequenceServer Cloud.
Accessing SWFSbase
SWFSbase is available SWFS members. Critically, access is restricted to supporting taxonomic identification in wildlife forensic casework and may not be used for research purposes.
To request access, you must be an SWFS member and complete the SWFSbase Terms of Use. Signed agreements should be emailed to: [email protected]. Please also include your full name, institution, and the email address you would like associated with your login.
The platform can be accessed here:
https://wildlifeforensics.sequenceserver.com
A brief user guide is available for download here.
Databases
SWFSbase contains the following BLAST databases:
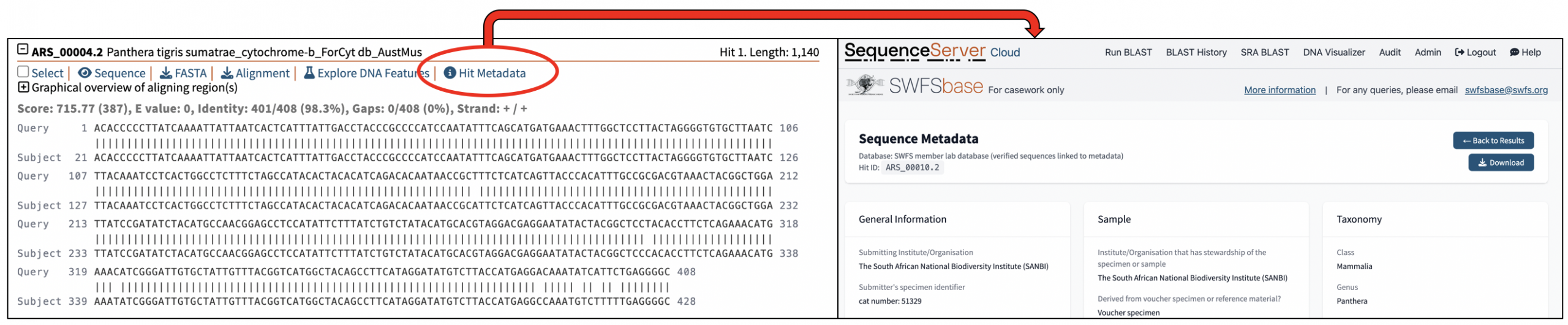
- SWFS member lab database (verified sequences linked to metadata): This is the key database within SWFSbase. It comprises vetted data contributed by multiple wildlife forensic laboratories, including data not available in other public repositories. Each sequence is linked to associated metadata to support assessment of specimen identification, sequence quality and reliability (see image below). Laboratories may contribute sequences by following the submission instructions provided below.
- Core nucleotide BLAST database: This is the entire NCBI core nucleotide database (“core_nt”), updated following NCBI's frequency (typically daily to fortnightly). These sequences are not curated within SWFSbase and are not linked to forensic metadata. The database is provided for convenience, allowing users to conduct searches without leaving the platform.
- Known erroneous NCBI sequences (from the NCBI core nucleotide database above): This is a small database that contains sequences from the NCBI core nucleotide database that have previously been identified as potentially erroneous. After conducting a search against the standard NCBI database, users may wish to query this database to determine whether top matches correspond to problematic records. To contribute to this database, please report any identified erroneous NCBI sequences to the SWFSbase team using the contact details provided on this page.
SWFSbase submission instructions
Laboratories that contribute data to SWFSbase play a vital role in enhancing the platform and furthering its mission. If you have highly reliable sequence data and associated metadata suitable for forensic analysis that you would like to contribute, please fill out this Google form.
The metadata template can be downloaded here, and instructions for the submission process can be accessed here.
SWFSbase contact information
SWFSbase subcommittee:
- Greta Frankham
- Kathy Moore
- Kelly Meiklejohn
- Kyle Ewart
- Marli de Bruyn
SWFSbase Director: Kyle Ewart
SWFSbase is hosted by SequenceServer Cloud, and is co-funded by SWFS, TRACE Wildlife Forensics Network, and ECO-SOLVE, a programme of the European Union.